Protein Design Competition — Community voting is now live!
This edition, we received over 10,000 proteins from over 600 designers. Head to proteinbase.com and vote for your favorite protein designers to have their sequences experimentally tested in the lab
The protein design competition community vote is now open!
This edition, we received over 10,000 proteins from over 600 designers. That’s more than 5x increase from our last competition. In other words, there’s a lot to explore!
How to vote:
➡️ Head to proteinbase.com
🧐 Browse the submissions
⭐ Star your favorites
You can vote for as many as you’d like.
The top submissions (up to 200 proteins in total) based on community votes will be experimentally tested in the lab.
Voting closes December 10th 11:59 PM PST.
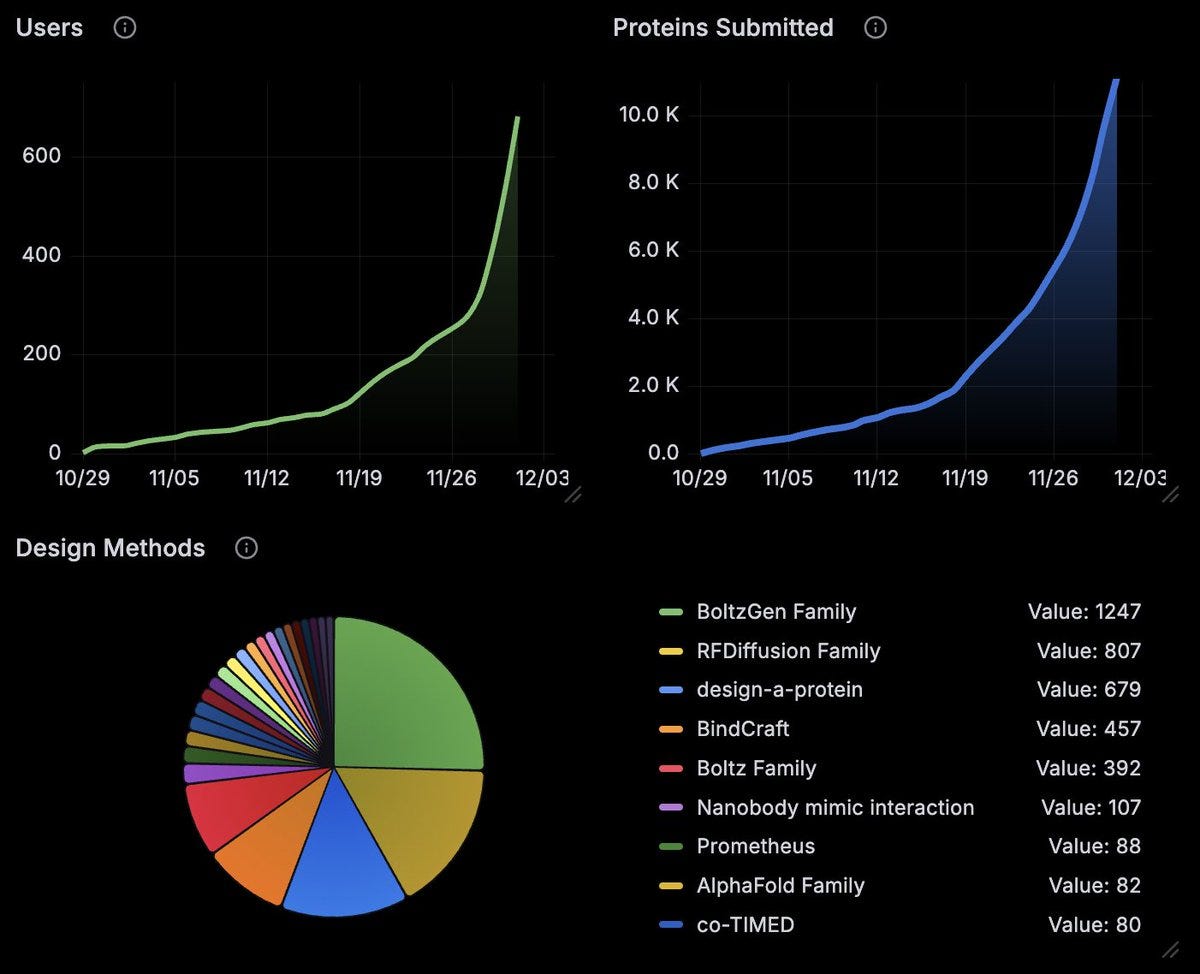
We’re hitting new records in terms of interest in protein design:
• Over 600 people participated
• Over 10’000 proteins were submitted
• Over 188 different protein design methods were used
When uploading their proteins, participants had the option to declare the design method used for each submission. Over half of all entries (5,648 proteins) included a self-declared method.
The most popular methods were:
• BoltzGen family: 1,247 proteins
• RFdiffusion family: 807 proteins
• Design-a-protein: 646 proteins
• BindCraft: 457 proteins
Overall, we saw impressive diversity and many new methods. We’re excited to see how they perform in the lab!
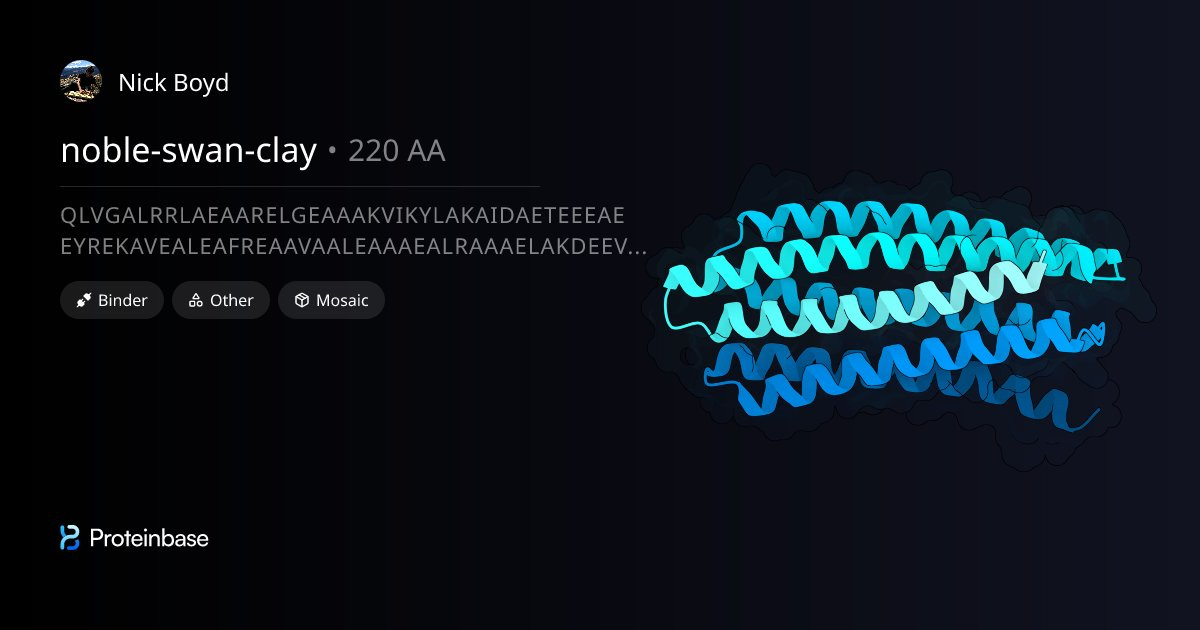
The Best ipSAE Award goes to Nick Boyd for his protein ‘noble-swan-clay’ with a whopping score of 0.92!
Check it out here
We received many questions about how ipSAE is calculated - we’ll share the exact methods and the reasoning behind in an email to participants soon
Want to dive deeper? Explore individual proteins, full design methods, and other stats in our Proteinbase collection:
https://proteinbase.com/collections/nipah-binder-competition-all-submissions